The following article contains a list of frequently asked question relating to views and displays in Scaffold PTM. For specific questions not covered in our documentation we are available by telephone Monday through Friday from 8 AM to 5 PM PST. Our toll free number is 1-800-944-6027. Additionally support can be contacted via email at support@proteomesoftware.com.
Views and Panes
What information can I see in the PTM List view?
This view contains a table listing the proteins and protein families included in the MS files loaded in Scaffold PTM. There are two sections for each protein after the name and accession number fields. The scores section displays the Scaffold protein probability and the sequence coverage. The modifications section gives information about modifications found on each protein and can be changed to view the number of modification sites, the number of unique modified peptides or the number of assigned modified spectra. These can be toggled using the Display Options dropdown menu. Clicking the header of any column will rearrange the data sorting it in ascending or descending order.
How is the Protein Sequence pane in Scaffold PTM different from others I've seen?
This tab shows an overall view of the best scoring PTM assignments highlighted along the selected protein sequence. Color coded schemes are used to convey information about the type of modification, its localization along the protein sequence and the degree of confidence associated to it.
What does the Peptide Score tab show me?
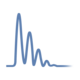
A 2D graph, showing the peptide ions score as a function of the peak depth & calculated using the Beausoleil algorithm, establishes the tolerated level of noise in the calculation of Ascore. The optimal peak depth used to calculate precise modification assignments is selected by looking at the earliest peak depth that represents the maximum difference between the two highest scoring site locations.
The legend identifies each curve with a corresponding peptide candidate that shows a particular combination of assigned modified amino acids.
How do I read the Spectrum & Ascore graph?
This tab shows the spectrum that identifies the peptide selected in the PTM Site pane. By hovering over this pane with the mouse and right-clicking it is possible to view the graph at different peak depths, allowing a better understanding of which peaks are considered in the Ascore calculation. Other viewing options usually available in Scaffold are also provided in Scaffold PTM.
In the lower part of the Spectrum & Ascore tab, there is a linear graph that shows highlighted in red and blue respectively, the y and b ions matched to site determining peaks for the selected peptide. These matching ions are used to calculate the Ascore for a particular modification.
Above each graph the modification identification and the calculated Ascore are listed along with the localization probability and the number of site determining peaks present in the spectrum over the total possible at a particular peak depth.
What are motifs? How can they help me?
Potentially significant modification trends that Scaffold PTM did not already list may become apparent on the Motifs Page. Scaffold PTM uses the Human Protein Resource Database in order to identify the enzymes that interact with the modification site associated with a motif. Then, the Motif Pane shows the likelihood that the identified motifs exist at each modification site across the experiment.
What does the Motif Representation Pane show me?
The Motifs Representation Pane creates a graphic representation of the probability that an amino acid might exist in the specific sites of a selected motif. Potentially significant modification trends that Scaffold PTM did not already list may become apparent here.
The visual representation centers the modification of the selected motif and displays the 12 flanking amino acids in the sequence, 6 to each side of the modification. Scaffold PTM scales each representative letter by the probability that it might exist in the flanking sequence of a motif. The amino acids in this pane are color-coded by chemical property.
Chemical Properties Color Code
| Chemical Property | Amino Acid | Color |
| Hydrophobic | AILMV | Black |
| Basic | HKR | Red |
| Special AA | GP | Green |
| Alcohol | ST | Light Blue |
| Cysteine | C | Yellow |
| Acidic | DE | Blue |
| Polar | NQ | Pink |
| Aromatic | FWY | Purple |
What is the difference between the PTM Spectrum Counts tab and the PTM Quantitation tab?
The Spectrum Counting tab displays a standard method for determining PTM abundance, while the PTM Quantitation tab displays quantitative PTM ratio data derived from a SQML generated in Scaffold Q+ and loaded into PTM. The ScaffoldPTM User's Guide goes into more detail about quantitative experiments in PTM.
Protein and Modification Details
Why do spectra in Scaffold PTM appear to be labeled differently than in Scaffold?
Because of the differences in scopes and aims between the two programs, the labeling of spectra is performed with slightly different procedures. Scaffold PTM tends to emphasize the presence of peaks that would help calculate a more reliable Ascore. Scaffold instead searches for a larger number of peaks and assigns more labels even if at times they are less likely to be correct. This means that sometimes certain peaks might be labeled in Scaffold and not in Scaffold PTM and vice versa. This fact does not influence the outcome of either program.
I have outside information that implicates an enzyme in a modification, but I don't see it in the Motifs Pane. Can I add it myself?
Yes, Scaffold PTM allows for custom motifs. For instructions on how to create a motif, please see page 29 of the ScaffoldPTM User's Guide.